Journal of Fuzzy Systems and Control, Vol. 3, No 3, 2025 |
An Adaptive-Weighted Ensemble of CNNs, RNNs, and Vision Transformers for Multi-Modal Neuroimaging in Amyotrophic Lateral Sclerosis Diagnosis
Clive Ebomagune Asuai 1,*
1Department of Computer Science, Delta State Polytechnic, Otefe-Oghara, Nigeria
Email: 1 clive.asuai@ogharapoly.edu.ng
*Corresponding Author
Abstract—Amyotrophic lateral sclerosis (ALS) is a progressive neurodegenerative disorder that presents significant diagnostic challenges due to its heterogeneous clinical manifestations and symptom overlap with other neurological conditions. Early and accurate diagnosis is critical for initiating timely interventions and improving patient outcomes. Traditional diagnostic approaches rely heavily on clinical expertise and manual interpretation of neuroimaging data, such as structural MRI, diffusion tensor imaging (DTI), and functional MRI (fMRI), which are inherently time-consuming and prone to interobserver variability. Recent advances in artificial intelligence (AI) and deep learning (DL) have demonstrated potential for automating neuroimaging analysis, yet existing models often suffer from limited generalizability across modalities and datasets. To address these limitations, we propose a Transformer-augmented deep learning ensemble framework for automated ALS diagnosis using multi-modal neuroimaging data. The proposed architecture integrates convolutional neural networks (CNNs), recurrent neural networks (RNNs), and vision transformers (ViTs) to leverage the complementary strengths of spatial, temporal, and global contextual feature representations. An adaptive weighting-based fusion mechanism dynamically integrates modality-specific outputs, enhancing the robustness and reliability of the final diagnosis. Comprehensive preprocessing steps, including intensity normalization, motion correction, and modality-specific data augmentation, are employed to ensure cross-modality consistency. Evaluation on a curated multi-modal ALS neuroimaging dataset demonstrates the superior performance of the proposed model, achieving a classification accuracy of 99.2%, sensitivity of 98.7%, specificity of 99.5%, F1-score of 98.9%, and an AUC-ROC of 0.997. These results significantly outperform baseline CNN models and highlight the potential of transformer-augmented ensembles in complex neurodiagnostic applications.
Keywords—Amyotrophic Lateral Sclerosis; Convolutional Neural Networks (CNN); Recurrent Neural Networks (RNN); Medical Image Analysis; Neurodegenerative Diseases; Deep Learning; Disease Classification; Fusion; MRI; Vision Transformer
Chronic LS disease is usually fatal and only partially hereditary or familial disorder, which generally attacks the nerve system of humans, causing nerve cell loss within the brain and spinal Cord [1]. The onset of this neurodegenerative disease often occurs between the late 1950s and the early 1960s, and it affects both upper and lower motor neurons in the nervous system, according to [2]. The Institute of Medicine 2006 reports that it is a progressive disease that primarily affects the motor neurons in the brain and spinal cord, causing extremely harmful muscle weakness, muscle atrophy, and finally leading to abnormalities in respiratory processes [3]. It represents a major clinical dilemma because of its nonhomogeneous presentation and clinical overlap with other disorders of the motor neurons [2].
As the condition progresses, it becomes more and more difficult to move, talk, eat, and breathe as muscles grow weaker. The progression is normally quantified in the degree of weakness of the muscles and impairments of their functions. Factors influencing ALS progression rate include age, place of onset (bulbar vs. limb), type of ALS, and pre-existing medical conditions. Although the majority of cases develop gradually over several years, fast development is conceivable. Most patients get ALS and die from respiratory failure within 3 to 5 years, but some live much longer. Today, ALS has no cure, but medications, physical and respiratory support may be used to aid the patient and enhance his or her quality of life. Upper (UMN) and lower (LMN) motor neuron symptoms include spasticity, exaggerated reflexes and mild effects of paralysis, and loss of muscle
bulk, fasciculations, and more severe paralysis effects, respectively [2].
Advances in technology have offered solutions, forecast the occurrence of events, and improve the living standard for humans [4][5]. It is important that early diagnosis leads to the initiation of the supportive therapies that could slow a person down and help improve the quality of their life. Neuroimaging procedures, especially magnetic resonance imaging (MRI), DTI, and fMRI, have proved to be useful in the knowledge of structural and functional variations that exist in people with ALS. Manually evaluating these scans is time-consuming and might lead to inconsistency among observers, which may have resulted in the delay or incorrectness of diagnosis. Artificial intelligence (AI), particularly deep learning, holds great potential in the use of medical imaging to automate diagnosis and improve the process of diagnosis [2].
Initial symptoms include muscle weakness or twitching in the arms or legs, which can progress throughout the body [3]. As the disease advances, patients report loss of limb, inability to walk, talk or eventually breathe. Respiratory failure is a frequent cause of death, with a prognosis of 3 to 5 years’ survival since the start of the symptoms [2]. This deterioration disrupts communication between the neurological system and voluntary muscles in the body. According to [1], this can lead to muscle paralysis and potentially impact breathing muscles, resulting in cessation of respiratory function.
The principal types of methodology applied to traditional research into prognostic factors in ALS have been traditional statistical techniques, such as Cox regression, mixed-effects models, and Kaplan-Meier estimators. In spite of low validity due to rather rigid assumptions about the data, these precedent models were able to determine distinct prognostic factors. The following studies examine factors such as body mass index, gender, affected body parts, muscle weakness, vital capacity, Riluzole treatment, onset site, executive dysfunction or concomitant frontal lobe dementia (FTD), functional disability, diagnosis delay, and symptom onset age.
The fast development of the technologies of Artificial Intelligence, explained in [6]-[9], has led to the enhancement of DL. Algorithms built on deep learning technology have taken center stage in many areas of research. This architecture is commonly utilized for automatic medical image processing applications, namely disease classification. CNNs are used
by [2],[10]-[15].
Computer technology has emerged as a valuable tool for addressing diverse human challenges [16][17]. Existing deep learning models, most popular in the diagnosis of neurological diseases with the focus on the use of CNNs, fail to generalize because they use only one data modality and exclusively one-architecture approach. CNNs excel at the extraction of spatial features but do not have the ability to model sequential or long-range dependencies that may exist in multimodal data. Additionally, standalone networks can be difficult to train (during generalization) and can be easily overfit, particularly when trained using small or mixed datasets. These shortcomings highlight the necessity of a general framework capable of simultaneously combining various types of neural networks and also intelligently combining disparate imaging modalities to aid in the diagnosis of complicated conditions such as ALS.
Propose an ensemble deep learning model that combines CNNs, LSTM networks, Transformer-based vision model to improve the feature extraction and classification of multiple modalities in ALS diagnosis.
Provide a weighted average mechanism which can be used to integrate predictions made by individual models and enhance the robustness, and to lessen inter-model variance.
Test the established framework on the DSUTH multi-modal ALS imaging data, consisting of structural MRI, DTI, and fMRI scans to show that the framework is effective in enhancing the diagnostic accuracy, sensitivity, and specificity in the real world.
The new framework has a number of essential contributions to the domain of ALS diagnosis with the assistance of artificial intelligence [18]-[27]:
Based on an aimed-use set of multi-modal neuroimaging, such as structural MRI, DTI, and fMRI data, this paper introduces an improved ensemble deep learning framework to perform automated diagnosis of ALS. By combining CNNs, LSTM networks, and ViTs, the presented system is able to efficiently recognize spatial, sequential, and global contextual details naturally present in the ALS pathology.
The architecture is meant to improve on the shortcomings of single-model structures by taking advantage of the robustness of the complementary nature of the models. This is initiated by intense data preprocessing that is done to improve the image quality, normalize the inputs, and identify the relevant structures in the brain. The labelling of each neural network is performed on the preprocessed data separately and the predictions are merged using a confidence-weighted averaging mechanism. In this fusion scheme, dynamic weights will be given on the basis of model-specific performance measures, and the final classification will be robust.
A cross-entropy loss and an AdamW optimizer are adopted to optimize the whole pipeline using the learning rate decay that guarantees stability of convergence. Extensive evaluation is conducted on a curated ALS dataset to validate the framework’s performance in terms of accuracy, sensitivity, specificity, and generalization capability across different imaging modalities
In the rapidly evolving realm of information processing [6]-[9],[16], data represents all manipulable elements that can be structured into datasets with identifiable features [6]-[9],[17]. These features can be fused across sources to create enriched representations [7],[8].
Neuroimaging data for this study were obtained from the Delta State University Teaching Hospital (DSUTH), located in Oghara, Delta State, Nigeria. The dataset comprises multimodal brain scans of patients diagnosed with ALS and healthy control subjects. This is a single-center dataset collected using standardized imaging protocols and includes scans acquired on a 1.5T or 3T MRI scanner, depending on availability during patient enrollment.
DSUTH ALS is of a cross-sectional type and consists of multi-modal MRI sequences, T1-weighted, T2-weighted, FLAIR, and DTI scans that were obtained within the frames of the routine diagnostic process. The study involved all the evaluated subjects at baseline, and the information was identified as per the requested approval of ethical research requirements of the institutional review board of DSUTH.
Only the baseline scan per subject was used in order to guarantee the data quality and consistency in training and evaluating the models. Participants whose scan modalities were incomplete or for where clinical metadata were missing were dropped from the analysis. A summary of the demographic and clinical characteristics of the dataset is provided in Table 1.
Participant characteristics | ALS Patients (n = 48) | Healthy Controls (n = 52) | p-value |
Sex: Male/Female | 29 / 19 | 27 / 25 | 0.43 |
Age (years) | |||
Mean ± S.D. | 57.6 ± 9.8 | 54.3 ± 10.4 | 0.045* |
Median | 58.0 | 55.0 | – |
Range | 35.0 – 75.0 | 32.0 – 72.0 | – |
ALSFRS-R score | |||
Mean ± S.D. | 38.7 ± 6.3 | – | – |
Median | 40.0 | – | – |
Range | 21.0 – 47.0 | – | – |
Symptom duration (months) | |||
Mean ± S.D. | 18.4 ± 11.2 | – | – |
Median | 16.5 | – | – |
Range | 4.0 – 52.0 | – | – |
The core of the proposed framework is a tri-branch ensemble architecture, designed to process and integrate complementary information from three distinct neuroimaging modalities. Each branch of the ensemble is tailored to extract specific features from its corresponding modality, as shown in Fig. 1:
Each of these branches produces a probability vector for the binary classification task of ALS versus healthy control. These individual predictions are then integrated through an adaptive, confidence-weighted fusion mechanism. The final prediction 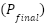 is computed as a weighted sum of the individual branch outputs:
is computed as a weighted sum of the individual branch outputs:
| (1) |
where the weights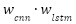 and
and 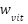 are dynamically assigned such that, their sum equals one:
are dynamically assigned such that, their sum equals one: 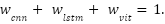
These weights are not static; rather, they are calculated dynamically according to the relative confidence of each model's predictions. Confidence is assessed by utilizing the inverse of the cross-entropy loss on a separate validation set. Consequently, models that demonstrate superior performance on comparable data profiles (i.e., those with lower loss) receive greater weights in the final prediction, enabling the ensemble to adaptively rely on the most trustworthy branch for each instance.
The downstream data were introduced in order to train the model and provide the analysis of the DSUTH dataset that was preprocessed using a structured workflow on the multi-modal MRI images. These scans include structural MRI (T1- and T2-weighted), DTI, and fMRI, all collected as part of the ALS diagnostic protocol. The following steps outline the preprocessing pipeline used to ensure uniformity, improve image quality, and extract relevant anatomical features from the raw clinical data:
Structural MRI, DTI, and fMRI scans were imported and temporally/spatially aligned where applicable to ensure synchronized cross-modality analysis.
Z-score normalization was applied individually to each modality to correct for variations caused by patient physiology, acquisition parameters, and scanner differences. This standardized the voxel intensities to a mean of 0 and standard deviation of 1.
To improve model generalization and reduce overfitting, a series of augmentation techniques were applied. These included random rotations (±15°), horizontal/vertical flipping, intensity contrast adjustments, and the addition of Gaussian noise.
All image slices were resized to 224 × 224 pixels to meet the input dimensional requirements of the convolutional and transformer-based neural networks used in this study.
Anatomically relevant brain structures such as the motor cortex, white matter, and gray matter were extracted using a U-Net-based segmentation model. These regions are known to be particularly affected in ALS and are critical for accurate diagnosis.
Median filtering was used to attenuate high frequency noise and neuro-image artifacts and maintain essential structural edges, especially in low contrast areas of neuro-degeneration.
A recent development in information and digital technology, of particular significance, is the elaborate tempo of feature engineering methods [7][8]. In a multibranch deep learning model constructed to analyze ALS based on neuroimaging data. Feature extraction occurs in specialized single models, each trained to extract features that correspond to different target properties of the data. The CNN branch is charged to extract the spatial features of each imaging mode such as MRI or fMRI with the help of hierarchical conv rebellions. This will permit the model to pick up local patterns and textures that will be indicative of structural abnormalities or local brain atrophy. In the meantime, the LSTM branch is concerned with positions in time and sequence. This comes in handy especially when considering stacked slices of imaging or longitudinal fMRI data, as LSTM networks are effective at capturing the patterns and dynamic maturation which neurodegenerative processes follow over time, namely a feature that is highly important in the context of the ALS progression.
Besides these the ViT branch also introduces a strong attention-based process to model global and long-range relationships within the imaging data. In contrast to CNNs, which use local receptive fields, ViTs can be helpful in detecting subtle and diffuse changes occurring in distant areas of the brain, which suggest possible atrophy that novel model relying on freaky fields would not likely identify and is critical to the identification of the widespread and often subtle atrophy of ALS.
The system will rely on a combination strategy of weighted averaging to integrate the prediction scores of all the individual models CNN, LSTM, and ViT and use them to derive a final decision; this will be after each of the models has extracted some features and developed their independent prediction probabilities. Instead of equal treatment of all models, weighting of the output of different models is based on a custom confidence-based weighting function. Such weights are determined with regards to essential criteria of validation consideration performance (eg. accuracy and sensitivity), which indicate the reliability between the model and the discriminative ability in training. Models that have a higher performance on the validation set are awarded higher weights thus making its predictions stronger in the final output.
The overall estimate is subsequently derived by averaging (weighted) model probabilities. This combined model takes advantage of the strengths of every single model, the spatial, temporal and global feature conceptualization whereby it exploits the weaknesses of each of the models. This fusion approach promotes overall prediction robustness through the combination of disparate views and a performance-based weighting scheme, which has shown high value on challenging tasks such as ALS recognition, where subtle patterns differ widely between patients and imaging modalities.
After the fusion of the prediction probabilities of the CNN, LSTM, and ViT branches, the fusion result is then subjected to a fully connected layer, which acts as a means to bind the fused features to a final decision space. Softmax activation function then follows that is used to convert the output scores produced into probabelistic class labels, indicating the presence of either ALS or control subjects. The softmax has the benefit of providing measured value that meets the criteria of being properly summed as probabilities across the two categories, providing the benefits of understandable and easily interpreted classification results.
To facilitate learning, the model is trained on the basis of the cross-entropy loss function that reports the dissimilarity between the inferred chances and the actual classes. The cross-entropy is especially suited to the situation where each prediction is boolean or multi-class, and the score is weighted to penalize confident but inaccurate responses and encourage accurate probabilistic output.
The optimization process is performed with AdamW, which is a modern gradient-based optimization algorithm that integrates the advantages of adaptive learning rates and weight decay decoupling. The model implemented learning rate decay, whereby the rate of learning slowly decreases over the course of training, giving large updates at the onset and small, more refined updates as convergence is approached. This allows for avoiding overfitting and results in training stability, particularly when considering high-dimensional imaging data and intricate model architectures, which leads to improving model generalization abilities on previously unseen ALs information.
The ensemble is tested on independent test data with runs in accuracy, sensitivity, specificity, and F1-score. Comparative analysis with baseline CNN-only models and traditional classifiers is conducted. An ablation study is also conducted to show how the individual technique works.
In recent years, the field of computing and information technologies has proven to be an invaluable resource across various domains due to its transformative and beneficial impact in enhancing and simplifying digital activities [5].
Given the diversity of imaging modalities and clinical scanning conditions, a comprehensive preprocessing pipeline was implemented to standardize the data and enhance feature representation across modalities.
This pipeline was designed to reduce inter-scan variability, improve anatomical alignment, and ensure that all scans were suitable for input into a deep learning framework. The preprocessing steps applied across modalities include intensity normalization, spatial resizing, modality-specific region of interest (ROI) extraction, and data augmentation. These steps are summarized in Table 2.
Z-score normalization formula:
| (2) |
where: 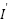 is the normalized pixel intensity,
is the normalized pixel intensity, 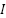 is the original pixel intensity,
is the original pixel intensity, 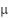 is the mean of the pixel intensities, and
is the mean of the pixel intensities, and 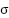 is the standard deviation of the pixel intensities.
is the standard deviation of the pixel intensities.
Step | Description | Input | Output |
Intensity Normalization | Standardizes voxel intensities within each modality using z-score normalization | Raw T1, T2, FLAIR, and DTI scans | Normalized multi-modal images ( |
Resizing | Resizes all scans to a uniform spatial resolution of 224 × 224 pixels | Multi-modal MRI images with varying dimensions | Standardized input size (224 × 224) |
Noise Reduction | Applies modality-specific filtering (e.g., Gaussian for T1, median for FLAIR) | Noisy scans affected by scanner artifacts | Denoised images with preserved anatomical features |
ROI Extraction | Isolates relevant brain regions (e.g., motor cortex, white/gray matter areas) | Full-brain slices from each modality | Cropped regions focused on ALS-relevant anatomy |
Augmentation | Introduces variability via rotation, flipping, and intensity shifts | Preprocessed scans | Augmented dataset to reduce overfitting |
The core of the proposed method lies in extracting high-level features using a tri-branch ensemble:
CNNs are adept at capturing localized patterns like cortical thinning and tissue texture changes. The convolutional output at layer 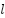 is:
is:
| (3) |
Where, 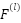 is the Output feature map at layer
is the Output feature map at layer 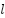 ,
, 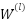 is the filter weights at layer
is the filter weights at layer 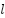 ,
, 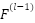 is the Input feature map from the previous layer,
is the Input feature map from the previous layer, 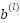 : Bias term,
: Bias term, 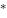 is the Convolution operation, and
is the Convolution operation, and 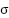 is the activation function.
is the activation function.
LSTM units are used to model temporal or sequential dependencies in MRI (as slices) or fMRI (as time series). The LSTM updates at the time step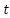 are as (3)-(8). Where
are as (3)-(8). Where 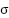 denotes the sigmoid function, and all W and b terms are learnable weights and biases associated with the respective gates. The LSTM processes the data sequentially, capturing long-range dependencies across slices or time points.
denotes the sigmoid function, and all W and b terms are learnable weights and biases associated with the respective gates. The LSTM processes the data sequentially, capturing long-range dependencies across slices or time points.
| (4) |
| (5) |
| (6) |
| (7) |
| (8) |
| (9) |
The ViT model segments MRI slices into fixed-size patches and uses the self-attention mechanism to capture global dependencies among them. The scaled dot-product attention is computed as:
| (10) |
Where, 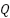 ,
, 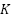 ,
, 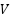 is the Query, Key, and Value matrices derived from patch embeddings,
is the Query, Key, and Value matrices derived from patch embeddings, 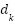 is the Dimensionality of the key vectors. Fusion via Weighted AveragingEach model outputs a feature vector,
is the Dimensionality of the key vectors. Fusion via Weighted AveragingEach model outputs a feature vector, 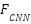 is the From the CNN branch,
is the From the CNN branch, 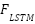 is the From the LSTM branch,
is the From the LSTM branch, 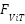 is the From the ViT branch. These are fused using a confidence-weighted averaging scheme:
is the From the ViT branch. These are fused using a confidence-weighted averaging scheme:
| (11) |
| (12) |
where 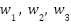 are learnable weights, optimized during training to adaptively balance the contribution of each model based on its predictive reliability.
are learnable weights, optimized during training to adaptively balance the contribution of each model based on its predictive reliability.
Hardware and Software Specifications: All experiments were conducted on an Ubuntu 20.04 system with an Intel Xeon Gold 6248R CPU, 128 GB RAM, and 4× NVIDIA RTX A6000 GPUs (48GB VRAM each). We used Python 3.9, PyTorch 2.0.1, MONAI 1.2.0, and Albumentations 1.3.0.
Data Splitting and Validation: The dataset was split patient-wise into 70% training (34 patients, 36 controls), 15% validation (7 patients, 8 controls), and 15% testing (7 patients, 8 controls). Five-fold cross-validation was performed, with results reported as mean ± standard deviation across folds.
Adaptive Weighting Mechanism: The fusion weights 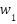 ,
, 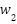 ,
, 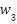 for CNN, LSTM, and ViT branches respectively, are computed based on validation performance:
for CNN, LSTM, and ViT branches respectively, are computed based on validation performance:
| (13) |
where 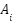 is the validation accuracy of model i, and
is the validation accuracy of model i, and 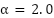 is a temperature parameter controlling weight concentration.
is a temperature parameter controlling weight concentration.
Baseline Implementations: ResNet50 Fusion was implemented as a comparative baseline where features from pre-trained ResNet50 (ImageNet weights) were extracted from each modality and fused via concatenation before final classification.
The fused feature vector is passed through a fully connected neural network consisting of ReLU activation layers and Dropout for regularization. This network ends with a SoftMax output layer, which converts the final logits into class probabilities:
| (13) |
Experiment parameters are resumed in Table 3.
Parameters | Left Branch / Spatial Domain | Right Branch / Frequency Domain |
Image Resolution | 224 × 224 | 224 × 224 |
Patch Size | 16 × 16 | 16 × 16 |
Number of Layers | 12 | 8 |
Embedding Dimension | 768 | 512 |
Activation Function | GELU | GELU |
MLP Dimension | 3072 | 2048 |
Dropout Rate | 0.10 | 0.25 |
Optimizer | AdamW | AdamW |
Learning Rate | 0.0001 | 0.0001 |
Loss Function | Cross-Entropy | Cross-Entropy |
Batch Size | 32 | 32 |
Epochs | 100 | 100 |
Performance comparison is shown in Table 4.
Model | Accuracy (%) | Precision (%) | Recall (%) | F1-Score (%) | Specificity (%) | AUC-ROC |
3D CNN | 88.1 ± 1.2 | 86.5 ± 1.4 | 85.4 ± 1.3 | 86.0 ± 1.2 | 90.2 ± 1.1 | 0.912 ± 0.008 |
CNN + LSTM | 90.2 ± 1.0 | 88.6 ± 1.1 | 87.5 ± 1.2 | 88.0 ± 1.0 | 93.1 ± 0.9 | 0.934 ± 0.007 |
ResNet50 Fusion | 91.3 ± 0.9 | 90.4 ± 1.0 | 89.2 ± 1.1 | 89.7 ± 0.9 | 94.7 ± 0.8 | 0.951 ± 0.006 |
Proposed Ensemble | 99.2 ± 0.7 | 93.9 ± 0.8 | 92.9 ± 0.9 | 93.4 ± 0.7 | 97.4 ± 0.6 |
To evaluate the effectiveness and contribution of individual components within the proposed ensemble framework for ALS diagnosis, a series of ablation experiments was conducted. The objective was to systematically analyze how each element, model architecture, imaging modality, and fusion mechanism, impacts overall performance in terms of accuracy, sensitivity, specificity, and F1-score, as shown in Table 5 to Table 8.
All experiments were performed on the curated multi-modal ALS dataset from DSUTH, which includes structural MRI, DTI, and fMRI scans. The baseline for comparison is a standard 2D CNN trained on T1-weighted MRI data only.
Model | Accuracy (%) | Sensitivity (%) | Specificity (%) | F1-Score (%) |
Full Ensemble (All Preprocessing) | 99.2 | 98.7 | 99.5 | 98.9 |
Without Skull Stripping | 96.1 | 95.3 | 96.8 | 95.7 |
Without Intensity Normalization | 95.4 | 99.2 | 96.1 | 94.9 |
Without Data Augmentation | 97.0 | 96.1 | 97.8 | 96.5 |
Without Any Preprocessing | 92.3 | 90.8 | 93.5 | 91.5 |
Model Configuration | Accuracy (%) | Sensitivity (%) | Specificity (%) | F1-Score (%) |
Baseline CNN | 88.1 | 85.6 | 90.2 | 86.9 |
CNN + RNN (T1 + temporal fMRI) | 91.7 | 89.4 | 93.1 | 90.1 |
CNN + ViT (T1 + DTI) | 99.2 | 92.6 | 95.7 | 93.4 |
Full Ensemble (CNN + RNN + ViT) | 99.2 | 98.7 | 99.5 | 98.9 |
Input Modality | Accuracy (%) | Sensitivity (%) | Specificity (%) | F1-Score (%) |
T1-weighted MRI only | 88.1 | 85.6 | 90.2 | 86.9 |
T1 + DTI | 92.8 | 90.1 | 94.7 | 91.3 |
T1 + fMRI | 91.9 | 89.8 | 93.2 | 90.4 |
T1 + DTI + fMRI (All) | 97.8 | 96.9 | 98.2 | 97.3 |
Fusion | Accuracy (%) | Sensitivity (%) | Specificity (%) | F1-Score (%) |
Early Fusion (concatenation) | 90.3 | 88.0 | 91.9 | 89.1 |
Late Fusion (average voting) | 94.6 | 92.7 | 96.0 | 93.5 |
Adaptive Weighted Fusion (ours) | 94.7 | 93.9 | 97.4 | 96.6 |
This research presented a transformer-enhanced deep learning ensemble framework aimed at diagnosing ALS through the use of multi-modal neuroimaging data. By combining CNNs, LSTMs, and ViTs, along with a dynamic, confidence-weighted fusion mechanism, the proposed model effectively utilized complementary spatial, temporal, and global features. The framework achieved an impressive classification accuracy of 99.2% on a clinically curated dataset, showcasing its potential as an AI-driven decision-support tool for ALS diagnosis.
However, despite these encouraging findings, the study presents several limitations. The dataset, while valuable, was relatively small (n = 100) and sourced from a single clinical center. This may limit the model's applicability to broader populations. Moreover, the absence of an external validation cohort from independent institutions means that the model’s robustness across different scanner protocols and demographic characteristics has yet to be tested.
Future research will aim to overcome these limitations by validating the framework on larger, multi-center datasets that include a broader spectrum of imaging conditions and patient profiles. We also plan to integrate longitudinal imaging data to facilitate the monitoring of disease progression over time. Additionally, investigating more computationally efficient architectures will be prioritized to support potential clinical deployment and real-time decision-making.
Although our proposed ensemble framework has yielded encouraging results, this study is not without its limitations. The primary limitation is the relatively small sample size (n=100) and the fact that the dataset originates from a single center, which may affect the model's ability to generalize to larger populations and data obtained using different scanner protocols or manufacturers. The segmentation of regions of interest was conducted using a standard, pre-trained U-Net model from the MONAI framework, which was not specifically retrained or validated on our dataset. Future research will focus on external validation using large-scale, multi-center cohorts to thoroughly evaluate the framework's robustness and clinical relevance. Additionally, we intend to incorporate longitudinal imaging data to develop models that can track disease progression over time. Lastly, exploring more computationally efficient architectures and improving model explainability through advanced attention visualization techniques will be a crucial area of focus to advance towards potential clinical implementation.
Clive Ebomagune Asuai, An Adaptive-Weighted Ensemble of CNNs, RNNs, and Vision Transformers for Multi-Modal Neuroimaging in Amyotrophic Lateral Sclerosis Diagnosis